Network Analysis with igraph
The igraph package in R is a
powerful tool for network analysis and visualization.
It provides a wide range of functions for creating, manipulating, and
analyzing graphs and networks.
This post showcases the key
features of igraph and provides a set of
graph examples using the package.
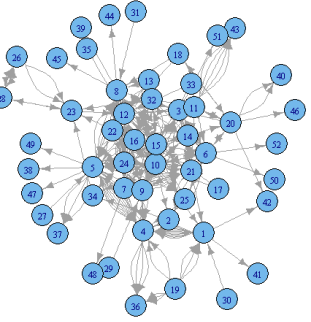
{igraph}